Copying data is wasteful, mutating data is dangerous
You have a large chunk of data—a NumPy array, or a Pandas DataFrame—and you need to do a series of operations on it. By default both libraries make copies of the data, which means you’re using even more RAM.
Both libraries do have APIs for modifying data in-place, but that can lead to other problems, including subtle bugs.
So what can you do?
In this article you’ll learn to recognize and apply the “hidden mutability” pattern, which offers a compromise between the two: the safe operation of copy-based APIs, with a somewhat reduced memory usage.
An example: using too much memory
Considering the following function:
def normalize(array: numpy.ndarray) -> numpy.ndarray:
"""
Takes a floating point array.
Returns a normalized array with values between 0 and 1.
"""
low = array.min()
high = array.max()
return (array - low) / (high - low)
If you call that function with an array whose values range from 30 to 60, 30 will become 0.0, 45 will become 0.5, and 60 will become 1.
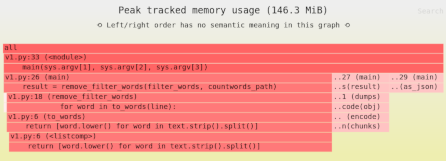
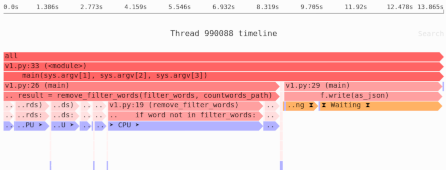
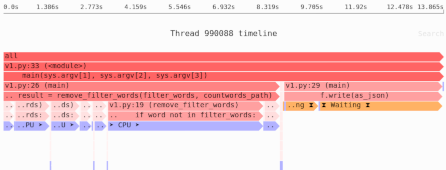
How much memory does this function use?
If the array uses A bytes, the function will use 3*A bytes of RAM:
- The original array, which is unmodified.
- The
array - lowtemporary array. - The result that gets returned from the function.
So how can we reduce memory usage?
In-place modification, aka mutation
To reduce memory usage, you can use in-place operations like += to do those operations on the original array:
def normalize_in_place(array: numpy.ndarray):
low = array.min()
high = array.max()
array -= low
array /= high - low
Similarly:
- Many NumPy APIs include an
outkeyword argument, allowing you to write the results to an existing array, often including the original one. - Pandas operations usually have an
inplacekeyword argument that modifies the object instead of returning a new one.
In all of these cases you’re “mutating” the data, modifying the original object.
And this saves memory!
In our example above, we’re using approximately A bytes of memory the whole time, as opposed to 3*A in the original version.
The problem with mutation
The problem with mutating data is that this can lead to unexpected behavior and bugs. Imagine if normalization was something you wanted to do in order to visualize your data:
def visualize(array: numpy.ndarray):
normalize_in_place(array)
plot_graph(array)
data = generate_data()
if DEBUG_MODE:
visualize(data)
do_something(data)
This code is buggy: do_something() likely expected the original data to be passed in, not the normalized data.
But depending on whether you’re in debug mode or not, do_something() will get called with different inputs.
More broadly, changing data out from under callers is not something that people using your code will expect—sometimes you’ll forget too, if enough time has passed.
So what should you do?
A flawed alternative: copy-before-call
You could require calling code to copy the array before calling visualize() if the intent is to preserve the original data:
data = generate_data()
if DEBUG_MODE:
visualize(data.copy())
do_something(data)
But that requires you and your colleagues to remember to do so every single time you call that function. Inevitably someone will forget and introduce a bug.
A better alternative: hidden mutability
The usual expectation you have when calling a function is that it does not mutate the inputs. But that doesn’t mean the function can’t use mutation internally, so long as it’s hidden from the outside world: mutation as an optimization, not an API choice.
Note: In an earlier version of this article I called this “interior mutability”, after a related concept from Rust, but some readers felt that was a distinct concept so I switched to “hidden mutability.” The Clojure programming language also has a similar concept.
Here’s what hidden mutability might look like in our case:
def normalize(array: numpy.ndarray) -> numpy.ndarray:
low = array.min()
high = array.max()
result = array.copy()
result -= low
result /= high - low
return result
From the caller’s perspective, this is the same as the original function: the input is never modified.
But we’ve reduced memory usage from 3*A to 2*A, since we don’t need to create a temporary array that is immediately thrown away.
Note: Whether or not any particular tool or technique will help depends on where the actual memory bottlenecks are in your software.
Need to identify the memory and performance bottlenecks in your own Python data processing code? Try the Sciagraph profiler, with support for profiling both in development and production macOS and Linux, and with built-in Jupyter support.
Explicit mutation is a last resort
Unnecessary data copying will waste memory, and once your data is big enough that will be a concern. But mutation is a cognitive burden: you need to think much harder about what your code is doing.
Luckily, quite often you’ll be able to use hidden mutability to reduce memory usage while still benefiting from the reduced cognitive overhead of immutable APIs. That means you should:
- Start out the easy way, by copying data.
- Next, optimize memory usage with the hidden mutability pattern.
- Finally, as a last resort expose mutation in your API.