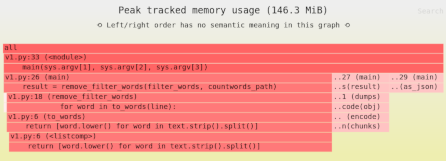
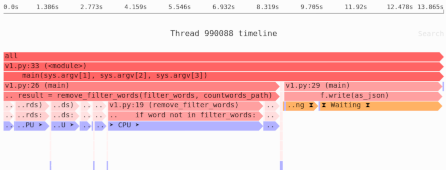
The mmap() copy-on-write trick: reducing memory usage of array copies
Let’s say you have an array, and you need to make some copies and modify those copies. Usually, memory usage scales with the number of copies: if your original array was 1GB of RAM, each copy will take 1GB of RAM. And that can add up.
But often, you’re just changing a small part of the array. Ideally, the memory cost would only be the parts of the copies that you changed.
As it turns out, there is an operating system facility that enables this: mmap()’s copy-on-write functionality.
In this article you will learn:
- How normal memory copies work.
- How to use
mmap()copy-on-write with NumPy. - How the underlying
mmap()copy-on-write mechanism works, and why it can be more efficient.
The problem with copying
If you want to modify a copy of an array, the normal approach is to allocate more memory and copy the contents of the original array into the new chunk of memory. For example:
>>> import numpy, psutil
>>> def memory_usage():
... current_process = psutil.Process()
... memory = current_process.memory_info().rss
... print(int(memory / (1024 * 1024)), "MB")
...
>>> array1 = numpy.ones((1024, 1024, 50))
>>> memory_usage()
428 MB
>>> array2 = array1.copy()
>>> memory_usage()
827 MB
In visual form, the allocated memory looks like this:
The pages are chunks of 4KB that are the unit of memory management for the operating system.
Saving memory with copy-on-write
In an ideal world, that second array would only store the differences from the first array: insofar as differences are few, the additional memory usage would be small.
And that’s where mmap()’s copy-on-write functionality comes in (or the equivalent API on Windows; NumPy wraps them both).
If you’re not familiar with mmap(), see my overview comparing mmap() with HDF5 and Zarr.
To use mmap() in this mode, we need a backing file.
While there is a file involved, so long as there’s enough memory available the file is almost an implementation detail; it needs to be there but it won’t impact performance much.
Note: On Linux you can go one step further and create an in-memory file using the
memfd_createAPI, which can be used in Python 3.8 and later by doingos.fdopen(os.memfd_create("mymemfile"), "rb+")and then “truncating” the file to be the appropriate size.
The numpy.lib.format.open_memmap() function will open a file of the appropriate size; we’ll start by creating our initial array:
>>> del array1, array2
>>> memory_usage()
20 MB
>>> open_memmap = numpy.lib.format.open_memmap
>>> mmap_array1 = open_memmap("/tmp/myarray", mode="w+", shape=(1024, 1024, 50))
>>> memory_usage()
22 MB
>>> mmap_array1[:] = 1
>>> mmap_array1[0] = 10
>>> memory_usage()
422 MB
Initially the array is just zeroes (at least on Linux and macOS; Windows may differ), so the operating system is clever enough not to allocate any new memory. Once we set some values, memory usage goes up accordingly.
Next, let’s create a copy: we’ll mmap() the same file with mode="c", which means copy-on-write.
On Unix systems like Linux or macOS, this translates to the MAP_PRIVATE flag to the mmap() API.
>>> mmap_array2 = open_memmap("/tmp/myarray", mode="c", shape=(1024, 1024, 50))
>>> mmap_array2[0, 0, 0]
10.0
>>> mmap_array2[10, 0, 1]
1.0
>>> memory_usage()
422 MB
We now have another copy of the array, with the same contents… but memory usage hasn’t changed!
Now let’s modify that second array, and we’ll see how memory usage goes up, but the original array is unchanged.
>>> mmap_array2[1:100] = 30
>>> memory_usage()
461 MB
>>> mmap_array1[1, 0, 0]
1.0
We have successfully made a copy of an array that:
- Doesn’t change the original array when mutated.
- Only stores those parts of the copy that have changed from the original, allowing us to save memory.
How copy-on-write works
When we mmap() a file with the MAP_PRIVATE flag, here’s what happens per the manpage:
MAP_PRIVATE
Create a private copy-on-write mapping. Updates to the
mapping are not visible to other processes mapping the same
file, and are not carried through to the underlying file. It
is unspecified whether changes made to the file after the
mmap() call are visible in the mapped region.
Notice that changes made to the file may or may not be visible, that behavior is unspecified. As a result, it’s best not to modify the original array.
Returning to our goal, we are saving memory by using copy-on-write. That means pages in the second array point to the first array until some change is made to them. Only when you write to the page does a copy get made and the writes applied.
Initially we mmap()ed /tmp/myarray with MAP_PRIVATE (by using mode="c"), and memory looked like this:
That is, we had another array, but no extra memory was used.
Then, we made some changes to part of the second array. Those pages that were modified get copied, and then modified—the rest still point to the original array. For example, if we modified some data in the first 4096 bytes in the array’s in-memory representation, a new page would be allocated that is a copy of the one in the first array:
Note: Whether or not any particular tool or technique will help depends on where the actual memory bottlenecks are in your software.
Need to identify the memory and performance bottlenecks in your own Python data processing code? Try the Sciagraph profiler, with support for profiling both in development and production macOS and Linux, and with built-in Jupyter support.
Paying only for what you change
The mmap() copy-on-write trick is useful when:
- You have a very large array.
- You are making copies and only partially modifying those copies.
In this situation, copy-on-write saves memory by only allocating memory for data that has actually changed. Just make sure not to modify the original array; you may have unexpected consequences depending on your operating system.
For other data structures, like dictionaries or lists, you can use immutable datastructures to reduce memory usage of mostly-similar copies; in Python the pyrsistent library is one implementation.