Reducing NumPy memory usage with lossless compression
If you’re running into memory issues because your NumPy arrays are too large, one of the basic approaches to reducing memory usage is compression. By changing how you represent your data, you can reduce memory usage and shrink your array’s footprint—often without changing the bulk of your code.
In this article we’ll cover:
- Reducing memory usage via smaller
dtypes. - Sparse arrays.
- Some situations where these solutions won’t work.
Using smaller dtypes
When you create an array in NumPy, it has a data type, a dtype that specifies what kind of array it is.
It might be an array of uint8 (unsigned 8-bit integers) or float64 (64-bit floating point numbers), and so on.
Different dtypes have different ranges of values they can represent:
- 16-bit uint range is 0-65535.
- 64-bit uint range is 0-18446744073709551615.
And they have different levels of memory usage; a 64-bit integer uses 4× memory than a 16-bit integer.
This gives us an opportunity to reduce memory usage: if your data is integers between 0 and 60K, there’s no point in using a 32-bit or 64-bit integer, you can use a 16-bit integer and use less memory.
>>> from numpy import ones
>>> int64arr = ones((1024, 1024), dtype=np.uint64)
>>> int16arr = ones((1024, 1024), dtype=np.uint16)
>>> int64arr.nbytes
8388608
>>> int16arr.nbytes
2097152
As you would expect, a 16-bit array uses 25% of the RAM that a 64-bit array does.
Sparse arrays
Whereas the dtype focuses on compression of individual cells in the array, sparse arrays focus on the overall structure of the array.
In particular, if your array is mostly zeros, why should you spend memory storing all those zeros?
A sparse array stores only the non-zero data, and all remaining data is assumed to be zero. There are different ways to implement sparseness, depending on the structure of your data, and we’ll focus on just one: coordinate-style.
Imagine a black and white picture of the stars: most of the background is black (i.e. zero), with occasional stars here and there. Instead of storing all the data, we can just say “at Y=123, X=500 there is a pixel with brightness 128”. Pixels that don’t get mentioned are assumed to have brightness 0. There is some overhead for recording X and Y for each star, but as long as most of the background is black this data structure will still save memory over the normal full array.
Example: reducing memory usage with a coordinate-style sparse array
In Python, the sparse library provides an implementation of sparse arrays that is compatible with NumPy arrays.
It mostly focuses on coordinate-style arrays, which it calls COO format.
Here’s an example based on one from the Sparse documentation: we create an 2D array with uniform noise between 0 and 1, and set 90% of the pixels to black. If you think of it as a picture, this is similar to the star example we gave above, lots of zeros with occasional bright spots.
We can then compare memory usage of the original and COO-sparse representation:
>>> import sparse, numpy as np
>>> arr = np.random.random((1024, 1024))
>>> arr[arr < 0.9] = 0
>>> sparse_arr = sparse.COO(arr)
>>> arr.nbytes
8388608
>>> sparse_arr.nbytes
2514648
The sparse array uses about 30% as much memory as the original array; only 10% of the array is non-zero, but there’s extra overhead from storing the X and Y coordinates.
Note: Whether or not any particular tool or technique will help depends on where the actual memory bottlenecks are in your software.
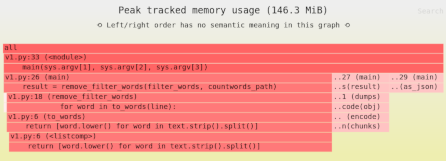
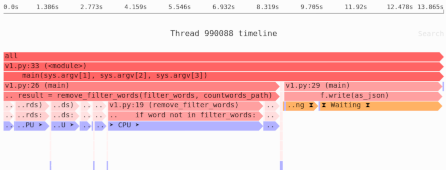
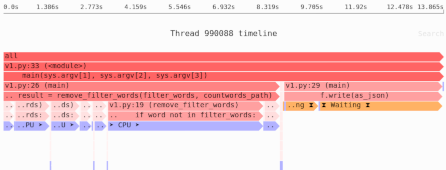
Need to identify the memory and performance bottlenecks in your own Python data processing code? Try the Sciagraph profiler, with support for profiling both in development and production macOS and Linux, and with built-in Jupyter support.
When these strategies won’t work
Let’s imagine you have an image stored in an array.
You realize that given the nature of the image, you can use a 16-bit unsigned integer dtype, and now you’ve limited the memory usage significantly.
Except—you want to use some functions from the excellent scikit-image library.
And the thing about scikit-image is that many of its functions will immediately convert the given array to a float64 dtype, if it isn’t already in that format.
So now you have the original 16-bit image and a new 64-bit image, for a total of 80 bits per pixel. In this situation you’re better off just storing the image as a 64-bit float in the first place, because at least 64-bit is better than 80-bit. Similar issues can apply to sparse arrays.
In short: even if you can find a smaller representation, the libraries you’re using might require a larger representation. Still, in many cases, just a tiny amount of changes to your code can reduce your memory usage, without changing your data at all.