Early speed optimizations aren’t premature
As you code, you might have a coworker, or friend, or a little voice in your head, reminding you of Knuth’s famous saying: “premature optimization is the root of all evil.” But what makes an optimization premature, anyway?
The short answer is that this aphorism is a tautology. “Premature” means “too early,” so we can rephrase the point as “doing things at the wrong time isn’t ideal.” Can’t argue with that!
The problem with this saying is that many people wrongly interpret it as “early optimization is the root of all evil.” In fact, writing fast software from the start can be hugely beneficial.
In order to reduce the scope a bit, I’m going to focus on one particular problem domain: data processing pipelines or batch jobs. This is the kind of software you often write when doing data science, or scientific computing: you load in some data, process them, spit out a result.
In this particular context, at least:
- Having sufficiently fast software from the start is a huge boost to productivity.
- Writing fast software doesn’t necessarily take any more work than writing slow software.
- Fast software is often just as good as slow software in terms of maintainability and correctness.
Note: The original version of this article mistakenly had a non-vectorized NumPy version that is not any faster, I’d copied in the wrong version of the code. Thanks to Gustav, and James Powell, for pointing it out.
Faster-running software means faster development
In “The counter-intuitive rise of Python in scientific computing”, the authors argue that the reason Python is becoming so popular for scientific computing is that it allows for easier exploration and experimentation when trying to solve a problem. If you don’t know the fastest algorithm or best solution in advance, iterating and experimenting can be much faster with Python, since it’s easy to write and modify.
This is a very good point, and on the face of it would seem to suggest that one should emphasize development speed, and that runtime speed is not important. After all, Python is a very slow language, or at least the default CPython interpreter is slow, and people use it anyway.
But this is only half the argument. The other half of the article’s argument points out that libraries like NumPy use vectorization to achieve much higher speeds, similar to compiled languages. You can also use just-in-time compilation with the alterative Python interpreter PyPy, or with Numba.
Runtime speed isn’t just important for the eventual real-world use of the code once development is over. If you’re working in this domain, it’s also critical to your productivity as a developer.
Vectorization in Python can give you a 25×-40× speedup over naive Python for loops.
If your job takes 1 minute in NumPy, it will take 30 minutes or more with plain old CPython.
That means going from being able to run your code 60 times an hour to only twice an hour, making it significantly harder to experiment and iterate on your code:
- Got a typo that breaks the program right at the end? It’ll be 30 minutes before you discover it.
- Is your latest idea a failure, or is there a bug in the implementation? You’re much more likely to dig in and discover it’s the latter if it only takes a minute to get results.
There are other programming languages that are as flexible as Python, and just as slow. But lacking equivalent technologies to NumPy or Numba, they are a bad choice for anything involving heavy computation or processing large amounts of data. Once a programming language is slow enough, it doesn’t matter how good it is otherwise: depending on your use cae, it might be non-starter.
In short, faster-running code can allow you to iterate and develop code faster. And that implies that writing your code to be fast from the start is a good investment.
But will it really? Won’t it take more time to write faster code, undoing the benefits of being able to iterate faster?
Writing faster software doesn’t always take more time
As we’ve seen, if you’re writing numeric array processing code with Python, you can use plain old Python, or you can use NumPy. For example, here’s a function written in plain old Python that takes a Python list and returns a Python list:
def normalize(values):
# Create a list of floating point numbers the same size
# as the input:
result = [0.0] * len(values)
max_val = max(values)
min_val = min(values)
for i in range(len(values)):
result[i] = (
(values[i] - min_val) / (max_val - min_val)
)
return result
And here is the same computation done with NumPy. Instead of accepting a Python list, it is designed to take a NumPy array and return a NumPy array.
import numpy as np
def normalize(arr):
max_value = np.max(arr)
min_value = np.min(arr)
result = arr.astype(np.float64)
result -= min_value
result /= (max_value - min_value)
return result
By using a better data representation and a faster implementation (aka vectorization), the NumPy version is 25× faster when run on the same, very large number of integers. That is a lot faster!
But writing the code with NumPy really doesn’t take any longer than writing the slow version.
It has the same number lines of code, the same basic algorithm: all it does is implement the for loop for you.
Which means that when it’s relevant, using NumPy is an optimization worth doing from the very start of coding. No need to write a vastly slower version first, just to avoid supposedly-premature optimization.
Is faster code less maintainable?
Compare the two code examples above, with and without NumPy.
They are both just as readable and maintainable. And NumPy is a well-supported, mature library; there’s no significant cost to adding it as a dependency.
Another example
Let’s look at a completely different example: you need to load 100MB of columnar data, composed of 100,000 rows, from an object store like AWS S3. Depending on how you choose to store the data, you could retrieve it using:
- 100,000 small 1K reads, sequentially.
- 100,000 small reads using parallelism: a pool of 100 clients, say. This will require a thread pool or an async runtime.
- One big 100MB read, which requires you to store the data in one big blob, using a suitable data format like Parquet.
If you assume a network latency of 1ms, elapsed time to retrieve the data will respectively be approximately:
- 100 seconds.
- 1 second.
- 1 second.
The third option is fast, simple, doesn’t require a thread pool or async runtime, and storing the data in that format may well be just as simple as storing 100,000 records. If you use AWS S3, it lets you run queries on CSV, JSON, and Parquet files it stores within a single blob, so you can still do subqueries if necessary elsewhere.
Why choose a slow design, or a fast but more complicated design, when you can choose a fast and simple design?
Note: Whether or not any particular tool or technique will speed things up depends on where the bottlenecks are in your software.
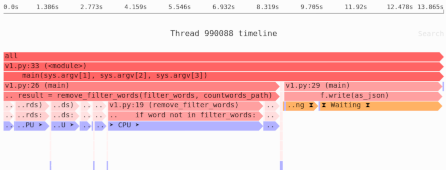
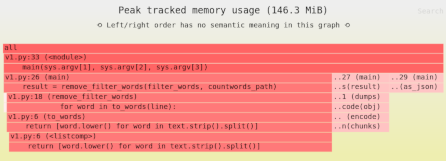
Need to identify the performance and memory bottlenecks in your own Python data processing code? Try the Sciagraph profiler, with support for profiling both in development and production on macOS and Linux, and with built-in Jupyter support.
If you’re not paying attention, your code is probably slower than it needs to be
At this point you could argue that these are just examples, hand-picked to make my point. This is quite true. One could find plenty of examples where writing faster code takes more effort, results in less maintainable code, or both.
In fact, beyond a certain point writing fast software is a form of transgressive programming: it requires breaking abstraction boundaries, which is not ideal. When the cost of performance involves significant time, or undermines code maintainability, and you don’t have an actual requirement for that speed (or evidence your changes will help), that’s where performance optimization can become premature.
That being said: there clearly exist situations where the fast version of your code is just as maintainable as the slow version, and just as easy to write. In the relevant domain, using NumPy is clearly better than not using NumPy.
If you aren’t in the habit of paying attention to speed, there’s a decent chance your code could have been written faster without additional work. The problems you’re solving likely aren’t unique. Which means someone has probably written a relevant library or tool, or documented an interaction pattern or algorithm, that will allow you to write faster code. Databases have indexes for a reason; programming languages provide multiple different data structures for a reason.
If your software runs faster, you can iterate, debug, and improve it more quickly. Not to mention that your users will be happier, and at scale you might also save some money and CO2 emissions.
So all else being equal, why wouldn’t you write faster software?