The limits of Python vectorization as a performance technique
Vectorization in Python, as implemented by NumPy, can give you faster operations by using fast, low-level code to operate on bulk data. And Pandas builds on NumPy to provide similarly fast functionality. But vectorization isn’t a magic bullet that will solve all your problems: sometimes it will come at the cost of higher memory usage, sometimes the operation you need isn’t supported, and sometimes it’s just not relevant.
For each problem, there are alternative solutions that can address the problem. In this article, we’ll:
- Recap why vectorization is useful.
- Go over the various limits and problems with vectorization.
- Consider some solutions to each problem: PyPy, Numba, and compiled extensions.
Vectorization has multiple meanings, we’re focusing on fast, low-level loops
To summarize a more detailed explanation of vectorization, there are three different meanings to the word in the context of Python:
- An API that operates on bulk data.
For example,
arr += 1will add 1 to every item in a NumPy array. - A fast API implemented in a low-level language (C, Rust), that operates quickly on bulk data. This will be our main focus in this article.
- Utilizing SIMD CPU instructions to speed up low-level operations even more. See the main writeup of vectorization for details.
We’ll be focusing on the second meaning, starting with the assumption the default Python interpreter, known as CPython.
When you run python on the command-line, you’re typically running CPython.
An example: adding an integer to a list of integers
Consider the following example:
from time import time
l = list(range(100_000_000))
start = time()
for i in range(len(l)):
l[i] += 17
print("Elapsed: ", time() - start)
A CPython list of Python integers is a series of pointers to fairly large objects, which from CPython’s perspective are just generic Python objects. Adding an integer to every item in a Python list involves:
- Looking up the type of each item in the list.
- Looking up the add function for that type.
- Calling that function with the original object and the Python object being added.
- Convert both of the Python objects into machine-code integers.
- Add the machine-code integers.
- Wrapping the resulting machine-code integer into a Python object.
- Looking up the next item in the Python list, which in itself is an expensive operation involving looking up the iterator methods for the range type, and then looking up the indexing operation for the list type, and then converting the index object into a machine-code integer…
That’s a lot of work!
In contrast, a NumPy array of integers is… an array of machine-code integers. Adding an integer to each item therefore involves only a handful of CPU instructions per item, it’s no different than the equivalent C code once the initial setup code is done.
Here’s the NumPy code:
from time import time
import numpy as np
l = np.array(range(100_000_000), dtype=np.uint64)
start = time()
l += 17
print("Elapsed: ", time() - start)
As you would expect, NumPy is faster. A lot faster.
| Implementation | Elapsed (secs) |
|---|---|
| CPython | 6.13 |
| NumPy | 0.07 |
Note: Whether or not any particular tool or technique will speed things up depends on where the bottlenecks are in your software.
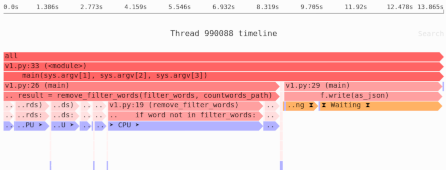
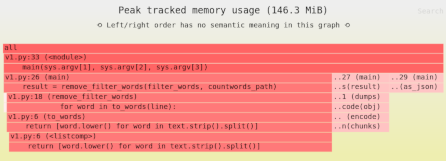
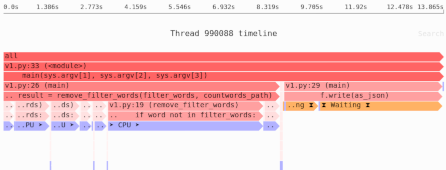
Need to identify the performance and memory bottlenecks in your own Python data processing code? Try the Sciagraph profiler, with support for profiling both in development and production on macOS and Linux, and with built-in Jupyter support.
The limits of vectorization
Vectorization makes your code faster, which is great… but it’s not a perfect solution. Let’s consider some of the problems.
Problem #1: Unnecessary large memory allocations
Let’s see you want to calculate an array’s items’ mean distance from 0. One way to do that is to calculate the absolute value of each item, then calculate the mean of the result:
import numpy as np
def mean_distance_from_zero(arr):
return np.abs(arr).mean()
That works, and the two calculations are both fast and vectorized… but this involves creating a whole new intermediate array to store the absolute values. If your original array is 1000MB, you’ve just allocated another 1000MB. You could overwrite the initial array to save memory, but you might need it for other reasons.
In practice, you can limit it somewhat with in-place operations so there’s only an extra copy, instead of N copies.
And there are libraries like numexpr and Dask Array that can do batching and smarter in-place updates for you, to reduce the memory burden.
Alternatively, you can just do the calculation item by item with a loop of your own… but that leads us to our second problem.
Problem #2: Only supported operations are fast
In order for vectorization to work, you need low-level machine code both driving the loop over your data, and running the actual operation.
Switching back to Python loops and functionality loses that speed.
For example, for our mean_distance_from_zero() function above, the obvious way to implement it without larger memory usage is like so:
def mean_distance_from_zero(arr):
total = 0
for i in range(len(arr))
total += abs(arr[i])
return total / len(arr)
But we’re back to doing a loop in Python, and the operations in Python; the code is slow again. That’s one reason why NumPy has to implement so much itself: any time you have to fallback to Python, things get slow. As an added penalty, in a compiled language there’s also the possibility that the compiler would find some way to optimize our second implementation, giving us a speedup that wouldn’t be possible with a series of separate vectorized operations.
Problem #3: Vectorization only speeds up bulk operations
Sometimes your code is slow because it’s doing the exact same operation on many data items of the same type. In that case, vectorization is your friend.
In other cases, your calculation is too slow but doesn’t match that particular pattern. At that point vectorization doesn’t help you, and you need some other solution to speed up your code.
Some solutions
There are three basic potential solutions we’ll cover here:
- A faster Python interpreter. PyPy is an alternative implementation of CPython which is much smarter: it can use just-in-time compilation to speed up execution by generating appropriate, specialized machine code.
- Generating machine code within CPython itself, by using Numba.
- Compiling custom code using Cython, C, C++, or Rust.
PyPy: a faster Python interpreter
If we have a Python list of Python integers, CPython (the default Python interpreter) doesn’t really do anything smart with it. That’s why we need to use NumPy arrays. PyPy is smarter; it can generate specialized code for that case. Going back to our original example:
from time import time
l = list(range(100_000_000))
start = time()
for i in range(len(l)):
l[i] += 17
print("Elapsed: ", time() - start)
We can compare CPython and PyPy:
$ python add_list.py
Elapsed: 6.125197410583496
$ pypy add_list.py
Elapsed: 0.11461925506591797
PyPy is almost as fast as the NumPy code, and … it’s just a normal Python loop. Since it knows you have a list of integers only, it can generate specialized, much-faster machine code for this specific case.
Unfortunately, PyPy doesn’t interface with NumPy particularly well; a naive non-vectorized loop over a NumPy array (e.g. the mean_distance_from_zero() we implemented above) is actually much slower in PyPy compared to CPython.
So if you’re trying to deal with missing functionality in NumPy, PyPy isn’t helpful; it even makes the slow unvectorized loop solution even slower.
If you’re dealing with code using standard Python objects, however, PyPy can be much much faster. So it’s best suited for speeding up cases where we’re not using NumPy or Pandas at all.
Numba: just-in-time functions that work with NumPy
Numba also does just-in-time compilation, but unlike PyPy it acts as an add-on to the standard CPython interpreter—and it is designed to work with NumPy. Let’s see how it solves our problems:
Extending NumPy with Numba
Missing operations are not a problem with Numba; you can just write your own.
For example, let’s implement our mean_distance_from_zero() in pure Python, and in Numba:
from time import time
import numpy as np
from numba import njit
# Pure Python version:
def mean_distance_from_zero(arr):
total = 0
for i in range(len(arr))
total += abs(arr[i])
return total / len(arr)
# A fast, JITed version:
mean_distance_from_zero_numba = njit(
mean_distance_from_zero
)
arr = np.array(range(10_000_000), dtype=np.float64)
start = time()
mean_distance_from_zero(arr)
print("Elapsed CPython: ", time() - start)
for i in range(3):
start = time()
mean_distance_from_zero_numba(arr)
print("Elapsed Numba: ", time() - start)
The result:
$ python cpython_vs_numba.py
Elapsed CPython: 1.1473402976989746
Elapsed Numba: 0.1538538932800293
Elapsed Numba: 0.0057942867279052734
Elapsed Numba: 0.005782604217529297
Like PyPy, Numba is generating specialized machine code for this function, though unlike PyPy, it can only do so for a subset of the Python language. The first call to Numba is much slower because Numba has to compile some custom machine code; after that the calls are extra fast. And as expected, CPython is much slower than Numba.
Because we can write custom add-on functions for NumPy using Numba, missing operations aren’t a problem. This also means we’re not forced to create unnecessary temporary arrays because of the limits of available NumPy arrays, so Numba can help us solve the first two problems we saw.
Numba is less helpful with non-vectorized operations, since they are not part of the project’s goals.
Compiled code with Cython (or Rust, or C, or C++)
So far we’ve seen two examples of just-in-time compilation: a whole interpreter, or just functions. The other approach to faster code is to compile code ahead of time. Using Cython, or Rust, or C, you can:
- Write fast, additional operations for NumPy using Cython’s native NumPy support, Rust’s
rust-numpy, or just using the Python C API. NumPy itself is mostly written in C, and existing NumPy extensions are also written in other languages like Fortran or C++. - For non-vectorized use cases, you can again use these languages to write fast extensions for Python.
You will, however, have to add a bunch of configuration to your packaging or build setup to compile these extensions before Python runs. Cython does actually have an option to compile on import, but that makes distributing your software harder since it requires users to have a compiler installed. Numba does not have that limitations.
Choosing a solution
For the vectorized operation use case, PyPy is not going to help. Ongoing efforts like the HPy project might make it more useful in the future.
That leaves Numba and compiled extensions.
- Because you need to compile in advance, which requires a compiler and more complex packaging, Numba is usually easier to set up.
- Numba can build different versions of the code for different types of integers or floats, with no extra work; with compiled code you have to compile a version for each, and perhaps write code for each as well. With Rust or C++ you can get around the need for code duplication with generics or templates.
- Numba is a very limited subset of Python, and debugging its limitations can be difficult. In contrast, compiled languages are much more flexible; Cython supports almost all of Python, Rust is a sophisticated language with excellent tooling, and so on.
For non-vectorized operations, PyPy will often make your code faster without any changes at all; you can just try you code and see if it’s faster. Otherwise, you’re left with optimization of existing code, or lacking that choosing the appropriate compiled language to write a compiled extensions.